CytoCensus
CytoCensus
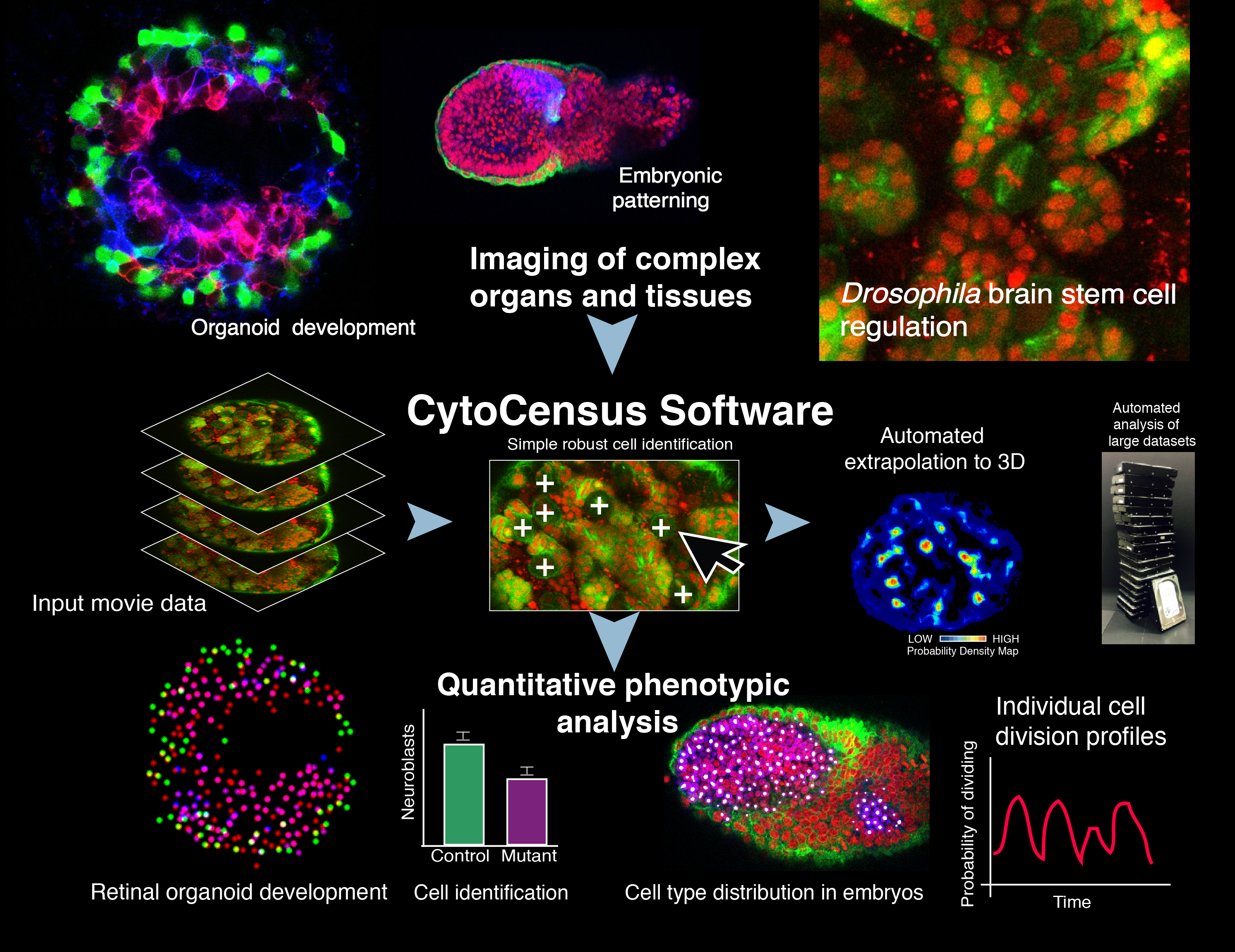
CytoCensus is a “point-and- click” supervised machine- learning image analysis software to quantitatively identify defined cell classes and divisions from large multidimensional data sets of complex tissues. In Hailstone et al. we demonstrate its utility in analysing challenging developmental phenotypes in living explanted Drosophila larval brains, mammalian embryos and zebrafish organoids. In comparative tests, a significant improvement in performance over existing easy-to-use image analysis software at cell detection.
CytoCensus is image analysis software intended to
allow biologists to identify and count cells and objects of interest
on large or numerous complex 3D microscopy images (e.g. time-lapse microscopy)
without detailed knowledge of image analysis
- Count cells/objects in 3D – if they are approximately round
- Use images of dense/complex tissue with low SNR
- Find cells of a particular size/colour/morphology
- Identify cell/object centres i.e. XYZ coordinates
- Determine counts and coordinates of cells in time-lapse image series
- Determine object counts in a user defined regions of interest (ROI) over time
- Compare the relative numbers of different object classes (or subclasses)
With a little FIJI/ImageJ knowledge, CytoCensus outputs can be used to:
- Determine average intensity across a class of cells
- Track cells that standard tracking struggles with (using CytoCensus probability maps and the ImageJ TrackMate plugin)
CytoCensus provides a simple interface for providing annotations and training.
1. Specify object size
2. Right-click and drag to define a region (red)
3. Left-click to annotate all the cells in the region (magenta).
4. When you're happy with it, Save the region (white)
5. Train the model and apply it to the rest of the image. (Add/adjust training if desired)
6. Find the peaks corresponding to your cells.
7. Apply the model to the rest of your images
A more detailed user manual is available for download with CytoCensus
Annotations
1) We ask the user to annotate a small number of cells (white crosses) within a series of regions in 2D.
Features + Training
2) We calculate a series of features that describe features of the image such as edges and shape.
3) We train a 'Random Forest'-like model that learns to associate these features with the proximity to the cell centres.
Proximity
4) We use this model to create a proximity map that represents how close each pixel is to the cell centre (red = close to centre, blue = far from centre)
5) Applying this to every plane in a 3D/4D image (by calculating image features and applying the model) we can create a 3D proximity map.
Detections
6) Using this proximity map, we apply a filter that finds 3D maxima: these are the detected cell centres
FAQs can be found on the Github site.
A user manual is bundled with the CytoCensus download:.
Found a bug? Have a suggestion? Raise an issue.